Select Publications
Computational epidemiologist. The projects below reflect three themes I work on: the epidemiology of zoonotic disease, infection biology in non-model species, and methods development for messy surveillance data.
Computational epidemiologist. The projects below reflect three themes I work on: the epidemiology of zoonotic disease, infection biology in non-model species, and methods development for messy surveillance data.





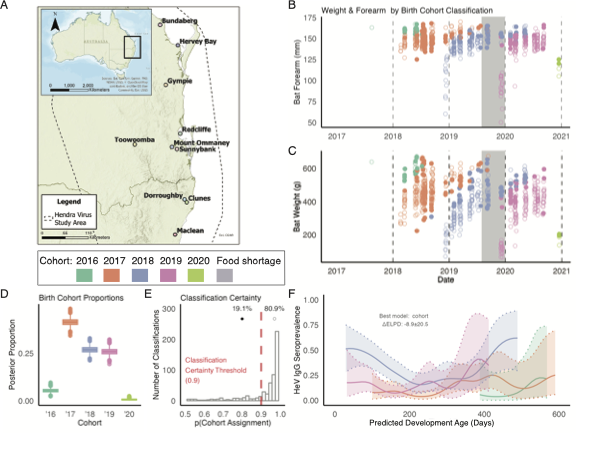
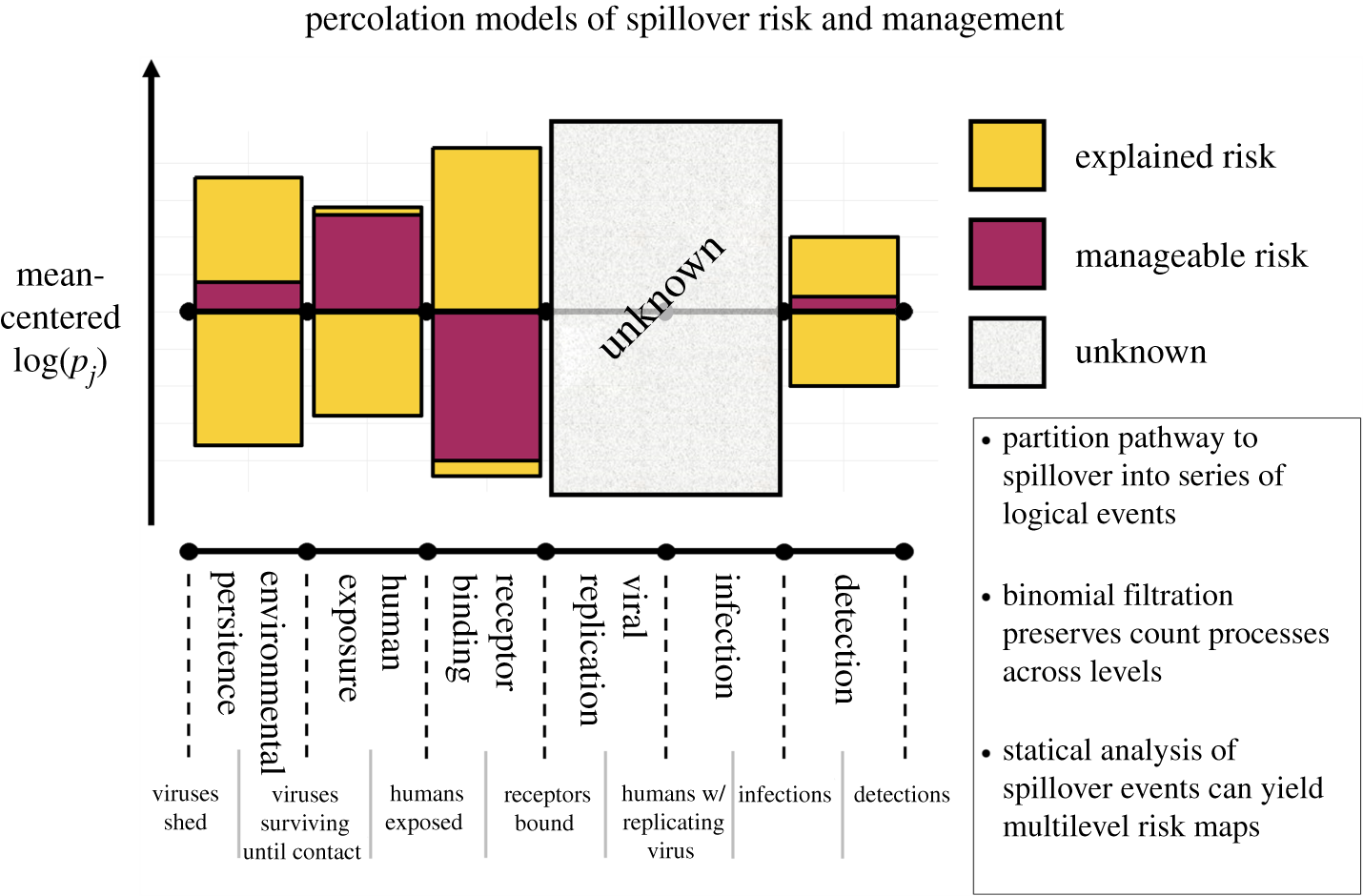
Predicting when and where pathogens will emerge means pulling signal out of long, noisy, often-pooled time-series. Whether the data are serological cohorts of wildlife, environmental samples, or clinical surveillance feeds, the underlying problem is the same: build models that flag high-risk windows before they become outbreaks. These papers develop and apply that toolkit.


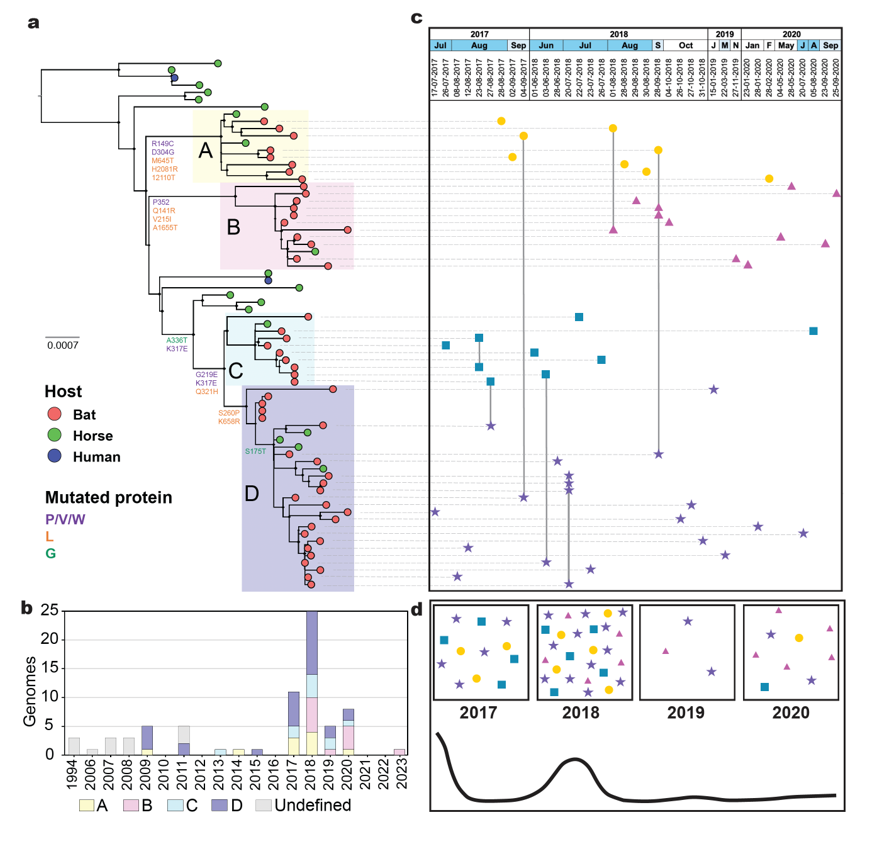
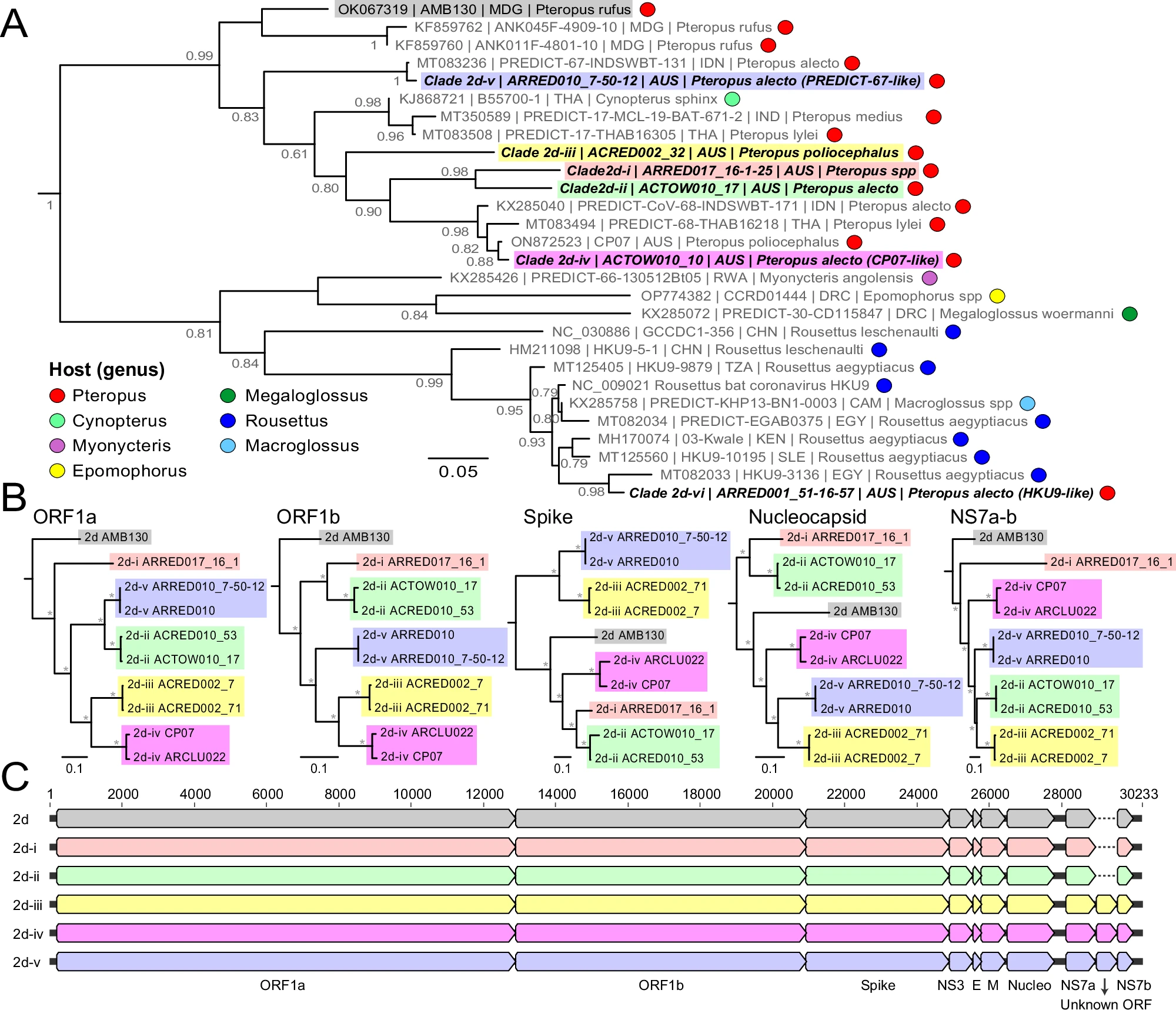

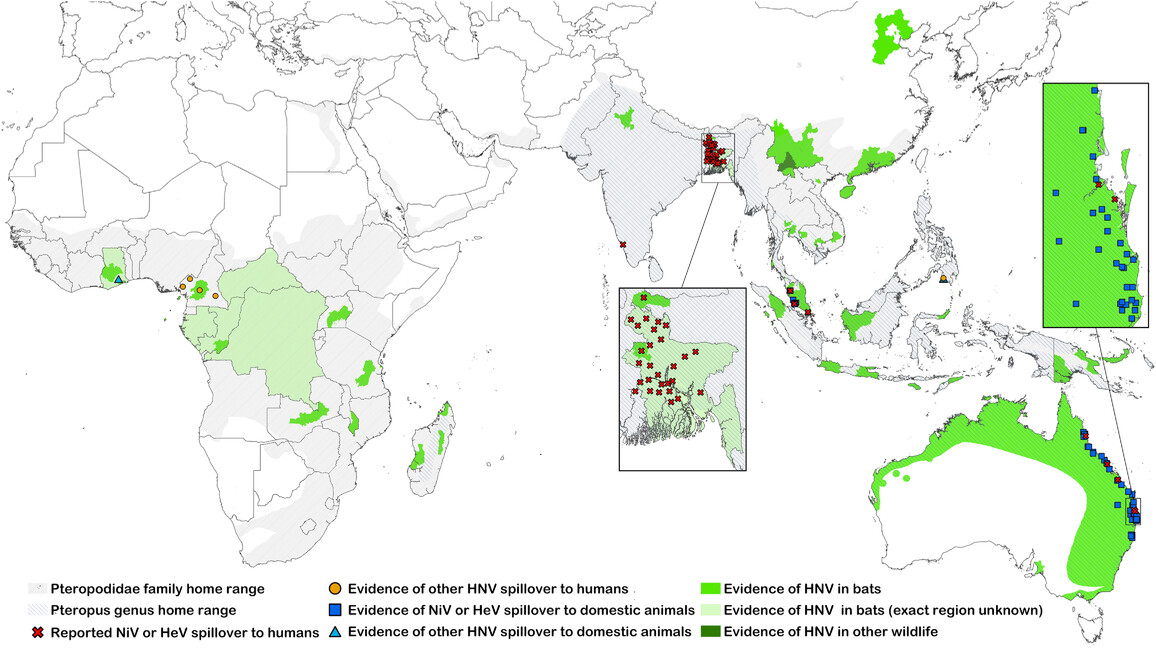

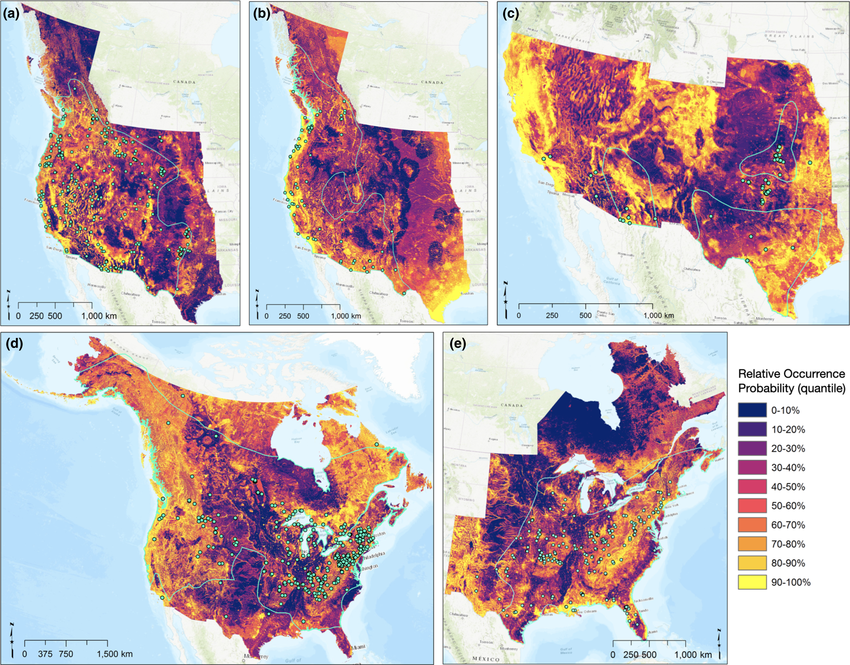
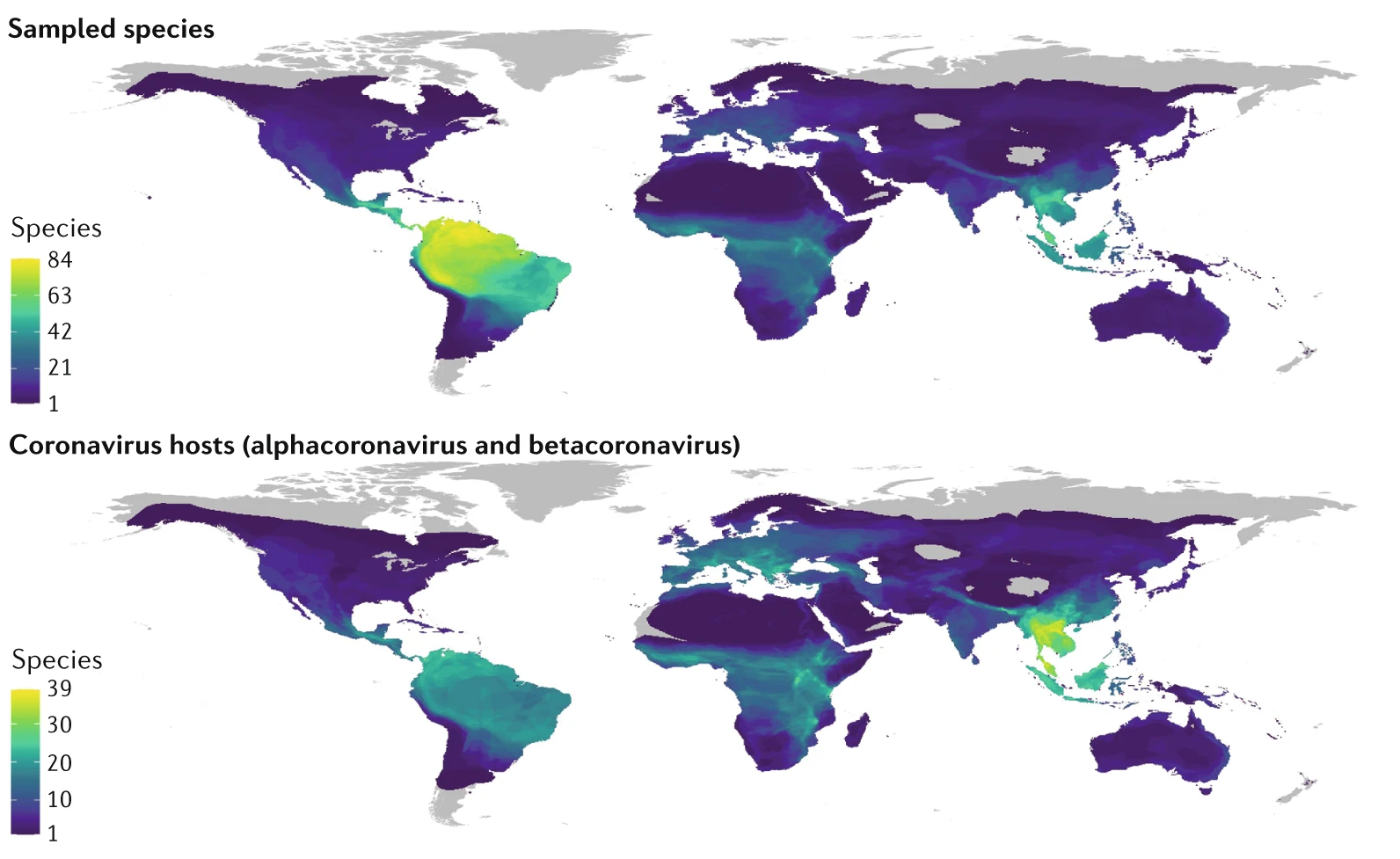
Where should we look for the next emerging pathogen? Bringing machine-learning, phylogenetic, and graph-based methods to large ecological and trait datasets — predicting reservoir hosts and prioritizing where surveillance investment is most likely to pay off.

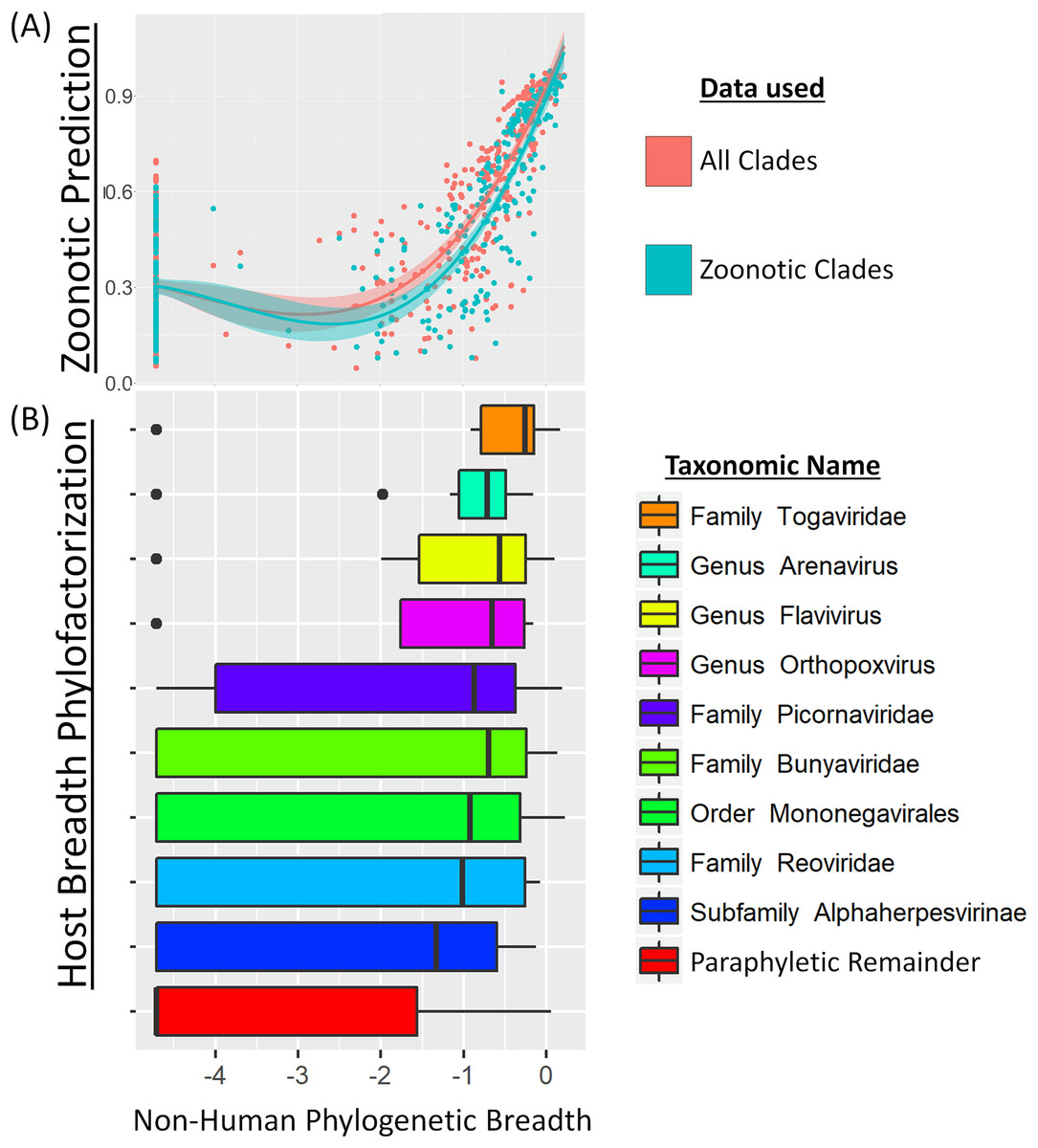

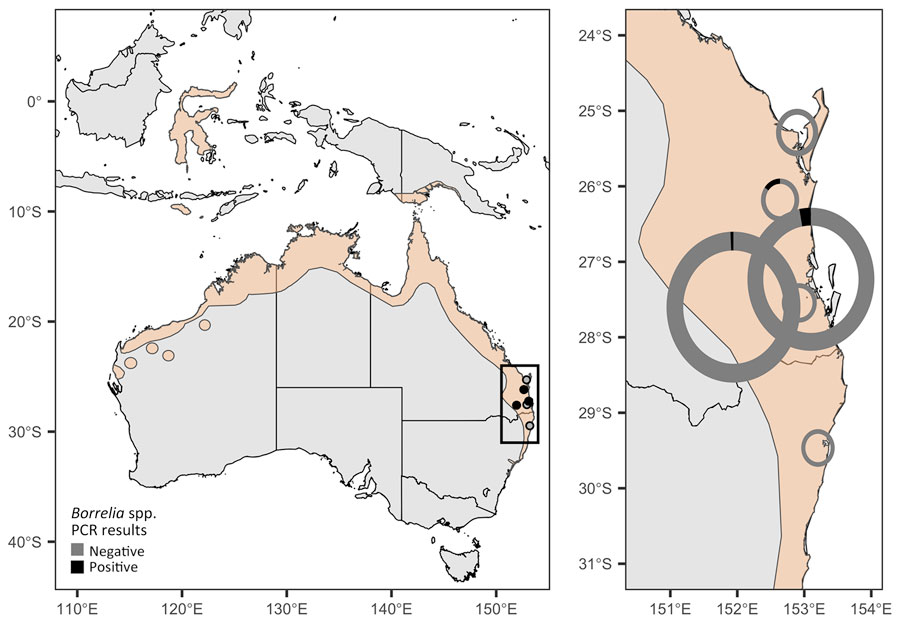
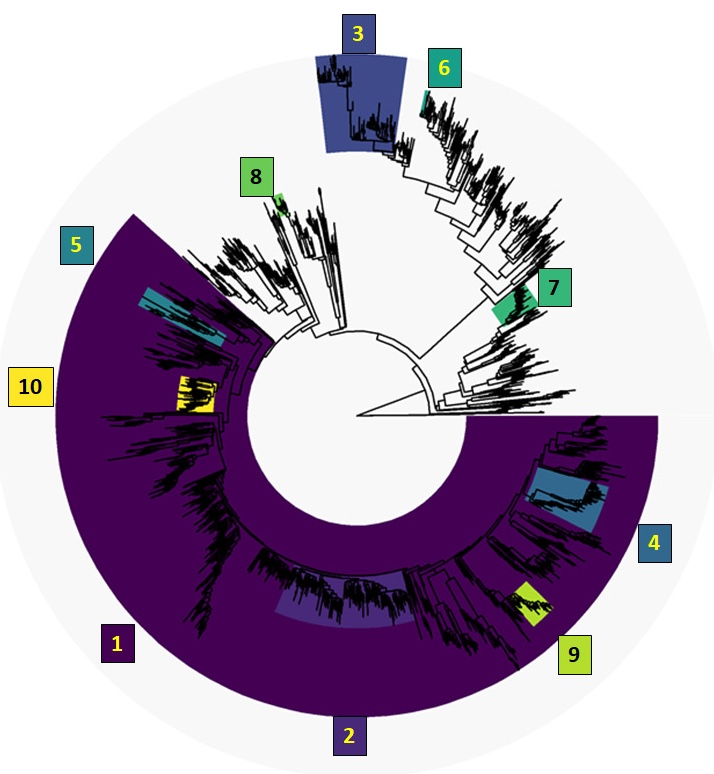
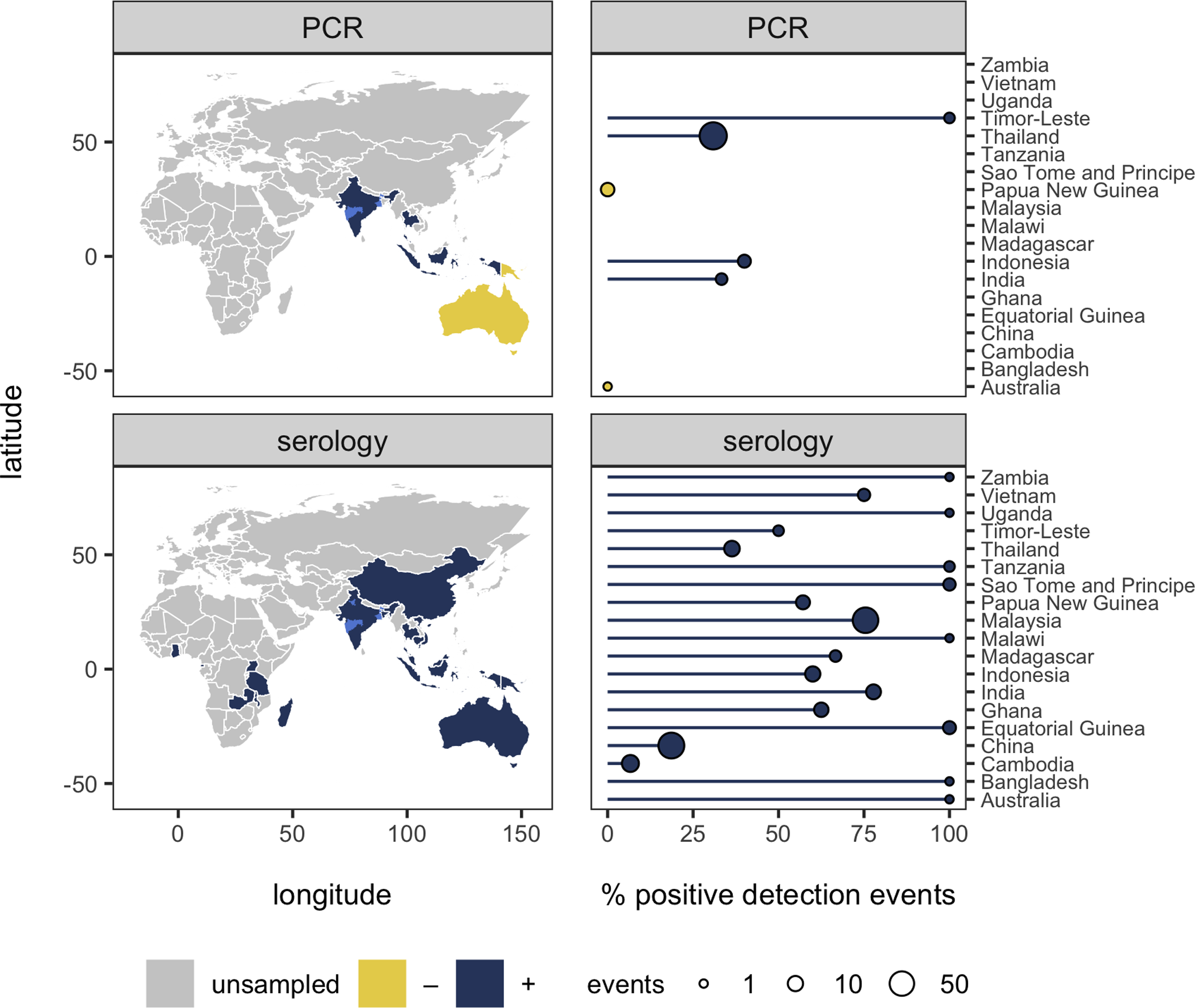
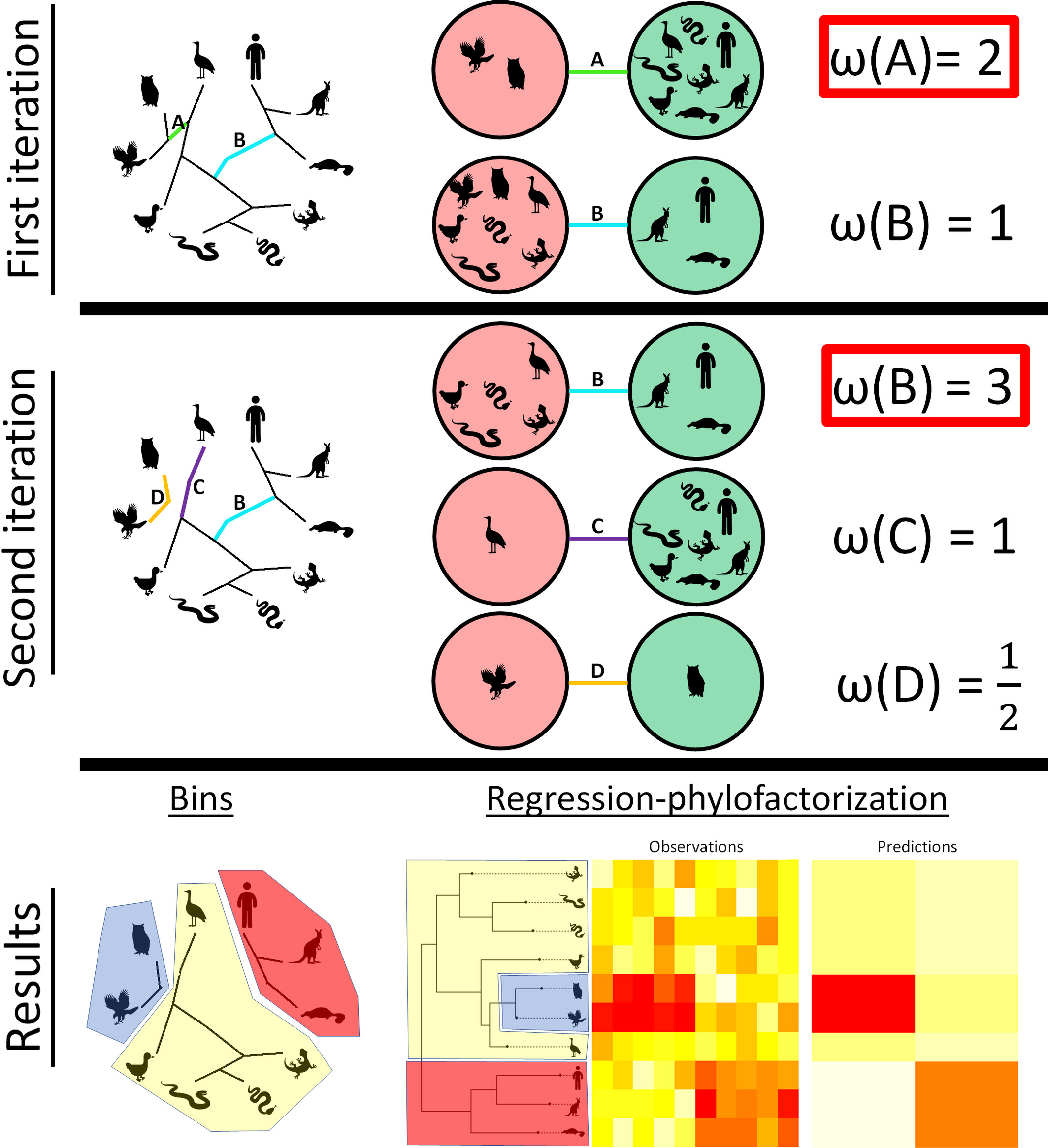
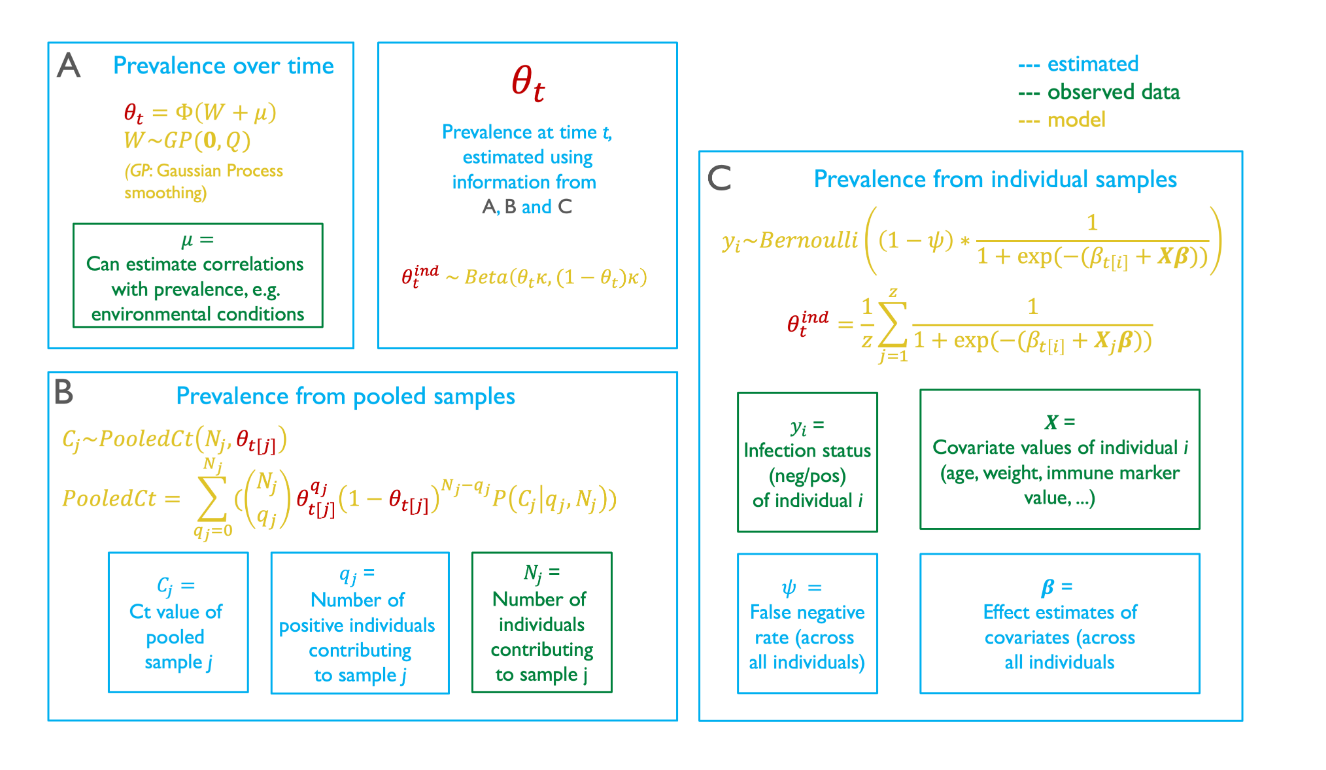
Real surveillance data is messy: pooled samples, sparse phylogenies, imperfect detection, heterogeneous platforms. Working with method-developers, I've contributed to tools that recover signal under those conditions — from phylogenetic graph-partitioning to Bayesian reconstruction of prevalence dynamics from pooled samples. The applications now extend well beyond what they were originally built for.
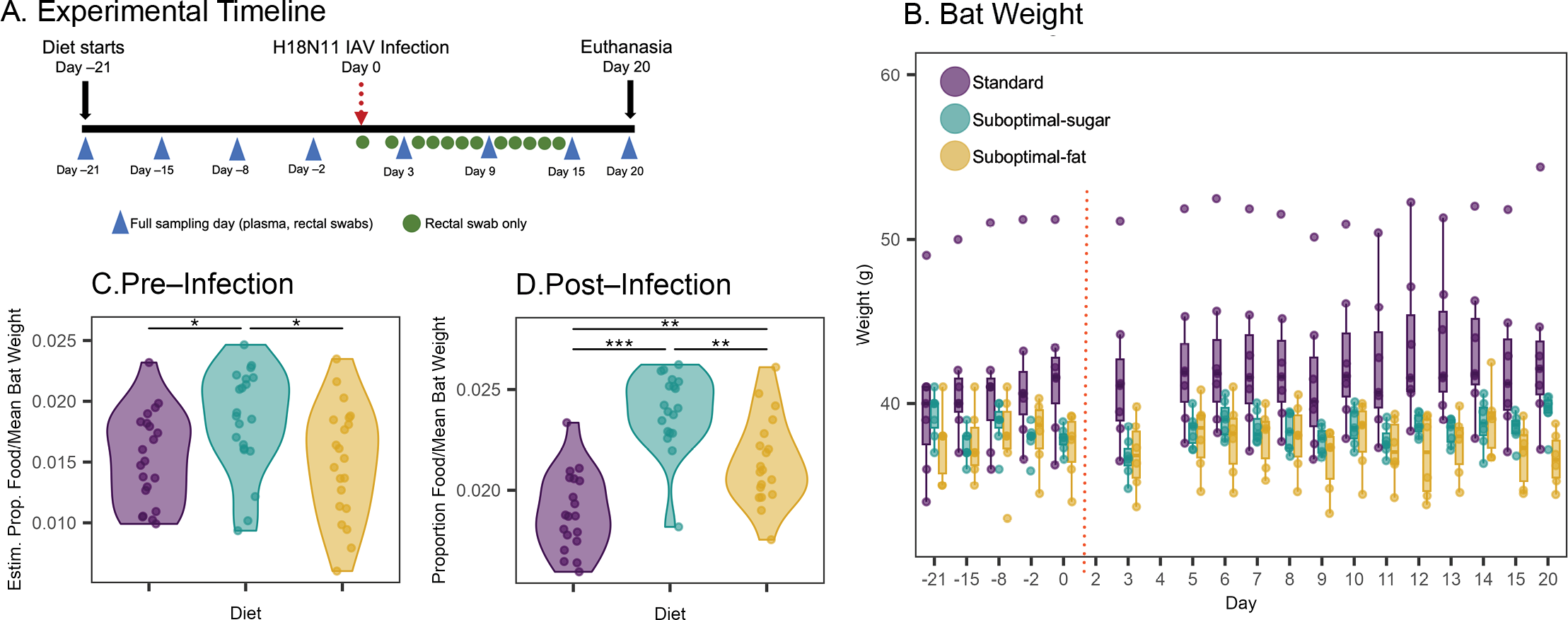
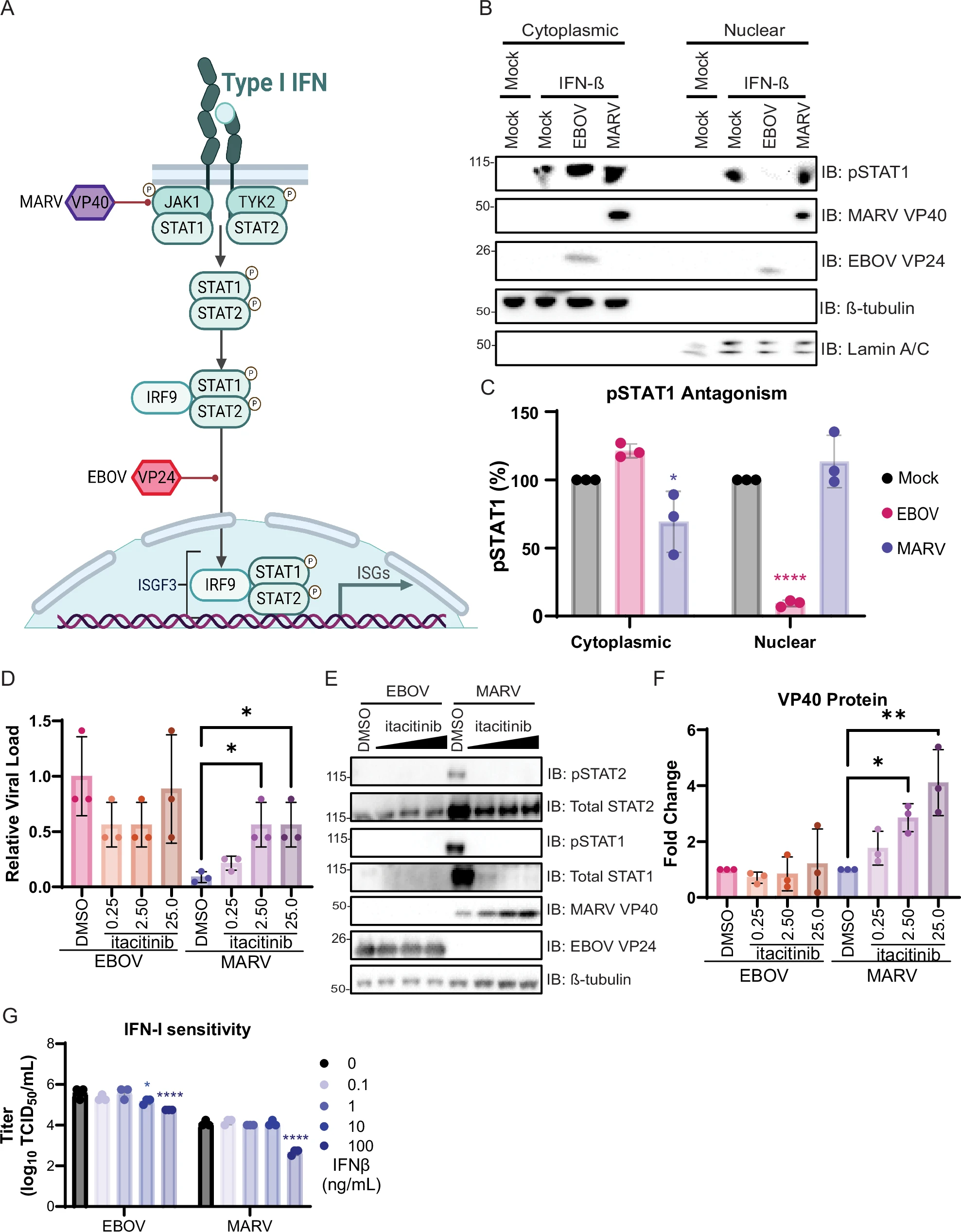
Most of what we know about mammalian immunity comes from a handful of inbred laboratory species. The pathogens that drive pandemics emerge from animals we barely understand. Captive colonies (two of which I established) let us run controlled infection experiments and apply Bayesian models to high-throughput sequencing data, mechanistically testing how non-model hosts tolerate dangerous infections and what conditions push shedding higher.


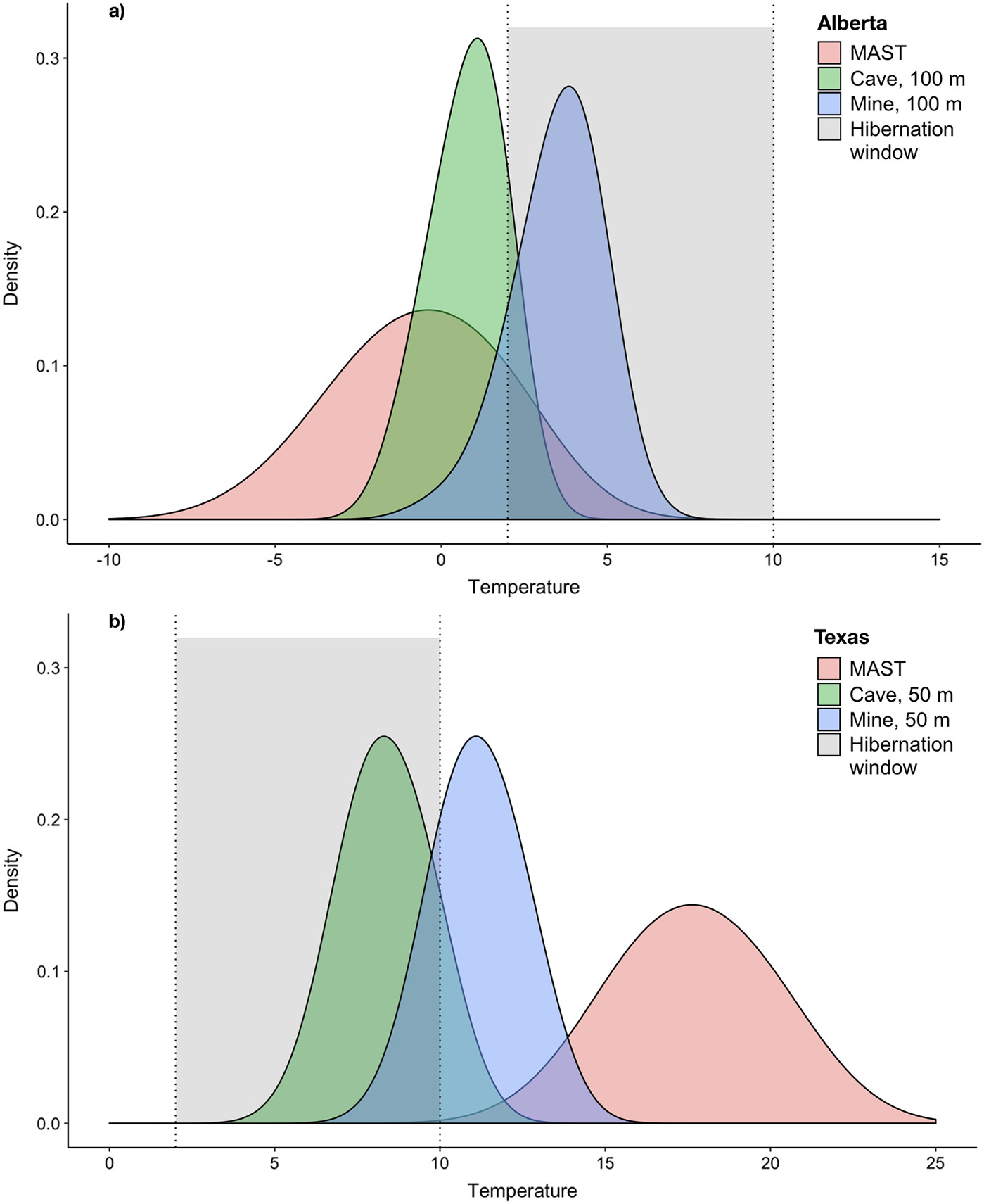
Endangered species share operational landscapes with the U.S. military, and disease — especially white-nose syndrome — compounds the problem. I built mechanistic and statistical models for DOD/DOE land managers, work that translated research into decision support and required ongoing technical communication with federal program officers.